Research: Difference between revisions
MarcRiedel (talk | contribs) Tag: visualeditor |
MarcRiedel (talk | contribs) |
||
| (41 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
''––'' '''George Bernard Shaw''' (1856 –1950) |
''––'' '''George Bernard Shaw''' (1856 –1950) |
||
Our research spans different disciplines ranging from digital circuit design, to algorithms, to mathematics, to synthetic biology. It tends to be ''inductive'' (as opposed to ''deductive'') and ''conceptual'' (as opposed to ''applied''). A recurring theme is building systems that compute in novel or unexpected ways with new and emerging technologies. |
Our research spans different disciplines ranging from digital circuit design, to algorithms, to mathematics, to synthetic biology. It tends to be ''inductive'' (as opposed to ''deductive'') and ''conceptual'' (as opposed to ''applied''). A recurring theme is building systems that compute in novel or unexpected ways with new and emerging technologies. |
||
==Storing Data with Molecules== |
==Storing Data with Molecules== |
||
''All new ideas pass through three stages:'' |
''All new ideas pass through three stages:'' |
||
| Line 11: | Line 11: | ||
''––'''''Arthur C. Clarke (1917–2008)''' |
''––'''''Arthur C. Clarke (1917–2008)''' |
||
Ever since Watson and Crick first described the molecular structure of DNA, its information-bearing potential has been apparent |
Ever since Watson and Crick first described the molecular structure of DNA, its information-bearing potential has been apparent. With each nucleotide in the sequence drawn from the four-valued alphabet of {A, T , C, G}, a molecule of DNA with ''n'' nucleotides stores ''2n'' bits of data. |
||
* Could we store data for our computer systems in DNA? ''"Can't be done '''–''''' ''too hard."'' |
|||
* Is it worth doing? ''"Definitely not. It will never work as well as our hard drives do."'' |
|||
* But one can store so much data so efficiently! "''I knew it was a good idea all along!''" |
|||
<br> |
|||
{| |
{| |
||
| |
| |
||
| Line 23: | Line 28: | ||
Peyton Okubo, John Stolzberg-Schray, Anil Reddy, and [http://mriedel.ece.umn.edu/wiki/index.php/Marc_Riedel Marc Riedel] |
Peyton Okubo, John Stolzberg-Schray, Anil Reddy, and [http://mriedel.ece.umn.edu/wiki/index.php/Marc_Riedel Marc Riedel] |
||
|- valign="top" |
|- valign="top" |
||
|''' |
|'''appeared in''': |
||
|[https:// |
|[https://pubs.rsc.org/en/content/articlepdf/2023/DD/D3DD00083D?page=search Royal Society of Chemistry – Digital Discovery], Vol. 2, pp. 1436–1451, 2023 |
||
|} |
|} |
||
| width="70" align="center" |<span class="plainlinks"> [https://mriedel.ece.umn.edu/wiki/images/8/8f/Manicka_Stephan_Chari_Mendonsa_Okubo_Stolzberg-Schray_Reddy_Riedel_Automated_Routing_of_Droplets_for_DNA_Storage_on_a_Digital_Microfluidics_Platform.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
| width="70" align="center" |<span class="plainlinks"> [https://mriedel.ece.umn.edu/wiki/images/8/8f/Manicka_Stephan_Chari_Mendonsa_Okubo_Stolzberg-Schray_Reddy_Riedel_Automated_Routing_of_Droplets_for_DNA_Storage_on_a_Digital_Microfluidics_Platform.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
[http://mriedel.ece.umn.edu/wiki/images/8/8f/Manicka_Stephan_Chari_Mendonsa_Okubo_Stolzberg-Schray_Reddy_Riedel_Automated_Routing_of_Droplets_for_DNA_Storage_on_a_Digital_Microfluidics_Platform.pdf |
[http://mriedel.ece.umn.edu/wiki/images/8/8f/Manicka_Stephan_Chari_Mendonsa_Okubo_Stolzberg-Schray_Reddy_Riedel_Automated_Routing_of_Droplets_for_DNA_Storage_on_a_Digital_Microfluidics_Platform.pdf Paper] |
||
| width="70" align="center" |<span class="plainlinks">[https://mriedel.ece.umn.edu/wiki/images/1/11/Dna-storage-and-computing.pptx http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
| width="70" align="center" |<span class="plainlinks">[https://mriedel.ece.umn.edu/wiki/images/1/11/Dna-storage-and-computing.pptx http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
||
[https://mriedel.ece.umn.edu/wiki/images/1/11/Dna-storage-and-computing.pptx Slides] |
[https://mriedel.ece.umn.edu/wiki/images/1/11/Dna-storage-and-computing.pptx Slides] |
||
|} |
|} |
||
<br> |
|||
[[File:Dna-computing-dalle.jpg|center|thumb|Storing data in DNA.]] |
|||
<br> |
|||
==Computing with Molecules== |
==Computing with Molecules== |
||
"''Biology is the most powerful technology ever created. DNA is software, protein are hardware, cells are factories''." |
|||
“''If I can’t create it, I don’t understand it.''” –– '''Richard Feynman''' (1918–1988) |
|||
––'''Arvind Gupta (1953– )''' |
|||
Over the past two decades, computing has moved from desktops and data centers into the wild. Embedded microchips – found in our gadgets, our tools, our buildings, our soils, and even our bodies – are transforming our lives. And yet, there are limits to where silicon can go and where it can compute effectively. It is a foreign object that requires a power source (a battery or an AC plug). We are studying novel types of computing systems that are not foreign, but rather an integral part of their physical and chemical environment: systems that compute directly with molecules. |
|||
Computing has escaped! It has gone from desktops and data centers into the wild. Embedded microcontrollers – found in our gadgets, our buildings, and even our bodies – are transforming our lives. And yet, there are limits to where silicon can go and where it can compute effectively. It is a foreign object that requires a electrical power source. |
|||
We are studying novel types of computing systems that are not foreign, but rather an integral part of their physical and chemical environments: systems that compute ''directly'' with molecules. A simple but radical idea: compute with '''acids''' and '''bases'''. An acidic solution corresponds to a "1" and a basic solution to "0". |
|||
<br> |
|||
{| |
{| |
||
| |
| |
||
| Line 53: | Line 66: | ||
|[https://www.nature.com/articles/s41467-023-36206-8 '''Nature Communications'''], Vol. 14, No. 496, 2023 |
|[https://www.nature.com/articles/s41467-023-36206-8 '''Nature Communications'''], Vol. 14, No. 496, 2023 |
||
|} |
|} |
||
|}The theory of [http://en.wikipedia.org/wiki/Chemical_kinetics mass-action kinetics] underpins our understanding of biological and chemical systems. It is a simple and elegant formalism: molecular reactions define ''rules'' according to which reactants form products; each rule fires at a ''rate'' that is proportional to the quantities of the corresponding reactants that are present. Just as electronic systems implement computation in terms of '''voltage''' (''energy per unit charge''), we can conceive of molecular systems that compute in terms of '''chemical concentrations''' (''molecules per unit volume''). We are studying techniques for implementing a variety of computational constructs with molecular reactions such as logic, memory, arithmetic, and signal processing. Although conceptual, we target [http://pubs.acs.org/doi/pdfplus/10.1021/ja906987s DNA Strand Displacement] as our experimental chassis. |
|||
<br> |
|||
{| align="center" |
|||
|| |
|||
[[Image:molecular-reactions-are-rules.gif|center|thumb|300px|Molecular reactions define ''rules'' according to which reactants form products. Here molecules of type A combine with molecules of type B to form molecules of type C, at a rate proportional to the molecular concentrations of A and B as well as a rate constant ''k''.]] |
|||
|| |
|||
|
|||
|| |
|||
[[Image:branch-migration.png|center|thumb|300px|We map abstract molecular reactions to DNA reactions. Through a process |
|||
called [http://pubs.acs.org/doi/pdfplus/10.1021/ja906987s DNA strand displacement], single strands of DNA displace parts of double strands, releasing other single strands.]] |
|||
|} |
|} |
||
<br> |
|||
[[File:Computing-with-chemistry.jpg|center|thumb|450x450px|Computing with Acids and Bases]] |
|||
'''Computational Constructs''' |
|||
<br> |
|||
We have developed a strategy for implementing digital logic with molecular reactions. Based on a bistable mechanism for representing bits, we implement a constituent set of logical components, including combinational components such as '''AND''', '''OR''', and '''XOR gates''', as well as sequential components such as '''D latches''' and '''D flip-flops'''. Using these components, we build full-fledged digital circuits such as a ''binary counters'' and ''linear feedback shift registers''. |
|||
It's more complex that acids and bases, but DNA is a terrific chassis for computing. We have developed "'''CS 101'''" algorithms with DNA: ''Sorting, Shifting'' and ''Searching'': |
|||
{| |
{| |
||
| |
| |
||
{| style="background:#F0E68C" |
{| style="background:#F0E68C" |
||
|- valign="top" |
|- valign="top" |
||
| width="100" | '''title''': |
| width="100" | '''title''': |
||
| width="500" | [https://mriedel.ece.umn.edu/wiki/images/b/b2/Solanki-chen-riedel-parallel-pairwise-operations-on-data-stored-in-dna-sorting-xor-shifting-and-searching.pdf Parallel Pairwise Operations on Data Stored in DNA: Sorting, XOR, Shifting, and Searching] |
|||
| width="500" | [[Media:Jiang_Riedel_Parhi_Digital_Logic_with_Molecular_Reactions.pdf | Digital Logic with Molecular Reactions]] |
|||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [[ |
| [[Arnav Solanki]], [[Tonglin Chen]], and [[Marc Riedel]] |
||
|- valign="top" |
|- valign="top" |
||
| '''appeared in''': |
|||
| [https://link.springer.com/article/10.1007/s11047-023-09964-z Natural Computing], 2023 |
|||
|- valign="top" |
|||
| '''presented at''': |
| '''presented at''': |
||
| [ |
| [https://drops.dagstuhl.de/opus/portals/lipics/index.php?semnr=16207 International Conference on DNA Computing and Molecular Programming], 2021 |
||
|} |
|} |
||
| |
| width="70" align="center" | |
||
<span class="plainlinks">[ |
<span class="plainlinks">[https://mriedel.ece.umn.edu/wiki/images/b/b2/Solanki-chen-riedel-parallel-pairwise-operations-on-data-stored-in-dna-sorting-xor-shifting-and-searching.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
<br>[https://mriedel.ece.umn.edu/wiki/images/b/b2/Solanki-chen-riedel-parallel-pairwise-operations-on-data-stored-in-dna-sorting-xor-shifting-and-searching.pdf Paper] |
|||
<br>[[Media:Jiang_Riedel_Parhi_Digital_Logic_with_Molecular_Reactions.pdf | Paper]] |
|||
| width="70" align="center" | |
|||
<span class="plainlinks">[http://mriedel.ece.umn.edu/wiki/images/d/d5/DNA27_Presentation.pdf http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/d/d5/DNA27_Presentation.pdf Slides] |
|||
|} |
|} |
||
Based on a bistable mechanism for representing bits, we have implemented logic gates such '''AND''', '''OR''', and '''XOR gates''', as well as sequential components such as '''latches''' and '''flip-flops''' with DNA. Using these components, we have built full-fledged digital circuits such as a ''binary counters'' and ''linear feedback shift registers''. |
|||
We have developed a strategy for implementing arithmetic with molecular reactions – operations such as '''increments & decrements''', '''multiplication''', '''logarithms''', and '''exponentiation'''. Try out our [http://mriedel.ece.umn.edu/chem-compiler/chem-compiler.pl compiler]: it translates arbitrary constructs from a '''C-like language''' into a robust implementation with molecular reactions. |
|||
{| |
{| |
||
| |
| |
||
{| style="background:#F0E68C" |
{| style="background:#F0E68C" |
||
|- valign="top" |
|- valign="top" |
||
| width="100" | '''title''': |
| width="100" | '''title''': |
||
| width="500" | [[Media: |
| width="500" | [[Media:Jiang_Riedel_Parhi_Digital_Logic_with_Molecular_Reactions.pdf | Digital Logic with Molecular Reactions]] |
||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [[ |
| [[Hua Jiang]], [[Marc Riedel]], [http://www.ece.umn.edu/users/parhi Keshab Parhi] |
||
|- valign="top" |
|- valign="top" |
||
| ''' |
| '''presented at''': |
||
| [http://www.iccad.com The International Conference on Computer-Aided Design], San Jose, CA, 2013. |
|||
| [http://dx.plos.org/10.1371/journal.pone.0021414 PLoS ONE], Vol. 6, No. 6, 2011.<br>[[Rate_Independent_Constructs_Supplementary_Information | Supplementary Information]] |
|||
|} |
|} |
||
| align="center" width="70" | |
| align="center" width="70" | |
||
<span class="plainlinks">[http://cctbio.ece.umn.edu/wiki/images/c/cf/Jiang_Riedel_Parhi_Digital_Logic_with_Molecular_Reactions.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<span class="plainlinks"> |
|||
<br>[[Media:Jiang_Riedel_Parhi_Digital_Logic_with_Molecular_Reactions.pdf | Paper]] |
|||
[http://mriedel.ece.umn.edu/wiki/images/d/db/Senum_Riedel_Rate-Independent_Constructs_for_Chemical_Computation.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/d/db/Senum_Riedel_Rate-Independent_Constructs_for_Chemical_Computation.pdf Paper] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks">[http://mriedel.ece.umn.edu/wiki/images/d/d2/Senum_Riedel_Rate-Independent_Modules_for_Chemical_Computation.pptx http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/d/d2/Senum_Riedel_Rate-Independent_Modules_for_Chemical_Computation.pptx Slides] |
|||
|} |
|} |
||
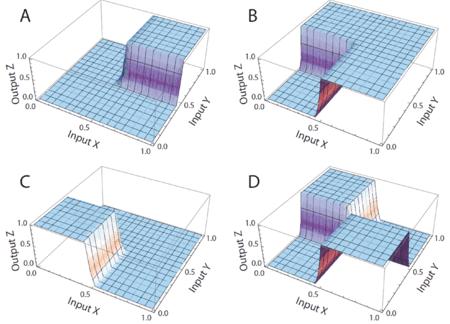
[[Image:dna-logic-gates.gif|center|thumb|450px| Simulations of DNA implementation of logic gates. The input signals are molecular concentrations X and Y; the output signal is a molecular concentration Z. (A) AND gate. (B) OR gate. (C) NOR gate. (D) XOR gate.]] |
|||
We have developed a strategy for implementing signal processing with molecular reactions including operations such as '''filtering'''. We have demonstrated robust designs for [http://en.wikipedia.org/wiki/Finite_impulse_response Finite-Impulse Response (FIR)], [http://en.wikipedia.org/wiki/Infinite_impulse_response Infinite-Impulse Response (IIR)] filters, and [http://en.wikipedia.org/wiki/Fast_Fourier_transform Fast Fourier Transforms (FFTs)]. |
|||
Also, we have performed signal processing including operations such as '''filtering''' and '''fast-fourier transforms (FFTs)''' with DNA. |
|||
{| |
{| |
||
| Line 146: | Line 152: | ||
<br> |
<br> |
||
{| align="center" |
|||
| |
|||
[[Image:dna-logic-gates.gif|center|thumb|450px| Simulations of DNA implementation of logic gates. The input signals are molecular concentrations X and Y; the output signal is a molecular concentration Z. (A) AND gate. (B) OR gate. (C) NOR gate. (D) XOR gate.]] |
|||
|| |
|||
|
|||
|| |
|||
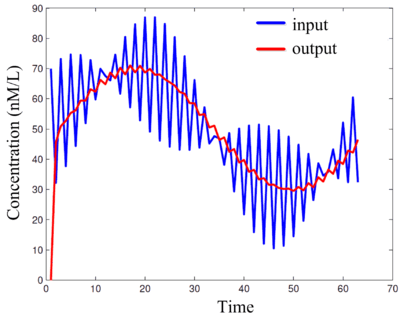
[[Image:moving-average-filter-simulation.gif|center|thumb|400px|Simulations of DNA implementation of a '''moving-average FIR filter'''. This filter removes the high-frequency component from an input signal, producing an output signal consisting of only the low-frequency component. Here the "signals" are molecular concentrations.]] |
[[Image:moving-average-filter-simulation.gif|center|thumb|400px|Simulations of DNA implementation of a '''moving-average FIR filter'''. This filter removes the high-frequency component from an input signal, producing an output signal consisting of only the low-frequency component. Here the "signals" are molecular concentrations.]] |
||
|| |
|||
|} |
|||
The impetus for this research is not computation ''per se''. Molecular computation will never compete with conventional computers made of [http://en.wikipedia.org/wiki/Integrated_circuit silicon integrated circuits] for tasks such as number crunching. Chemical systems are inherently slow and messy, taking minutes or even hours to finish, and producing fragmented results. Rather, the goal is to create “'''embedded controllers'''” – viruses and bacteria that are engineered to perform useful molecular computation ''in situ'' where it is needed, for instance for drug delivery and biochemical sensing. |
|||
[[Image:embedded-controller.png|center|thumb|350px|Molecular computation is applicable to the design of ''embedded controllers'': engineered bacteria and viruses for tasks such as drug delivery.]] |
|||
Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
||
| Line 164: | Line 160: | ||
==Computational Immunology== |
==Computational Immunology== |
||
“'' |
“''Physics is the study of the simple things in the Universe. Biology is the study of the complex ones.'' ” |
||
–– '''Richard Dawkins''' (1941– ) |
|||
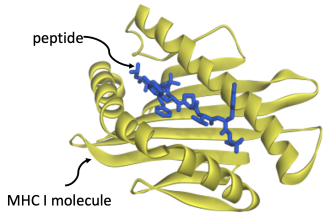
Cellular immunity allows circulating T-cells to kill off infected cells. When a cell is infected with a virus, it hijacks the host cell’s machinery, forcing it to make viral proteins. Our cells have a defense mechanism: they chop up such proteins into fragments, called peptides, and transport them to the cell surface, bound to MHC I molecules. Presented this way on the cell surface, T-cells can identify a cell as being infected and can destroy it using toxins. If this mechanism succeeds, an infection is stopped in its tracks: T-cells kill off infected cells before they can do damage. If it fails, then infected cells become factories for reproducing copies of the virus and full-blown disease results. |
|||
<br> |
|||
We are studying a problem that computer science currently judges to be very difficult: '''''predicting cellular immunity'''''. It centers on the question of how strongly molecules binds to one another. The molecules in question are ''peptides'' – fragments of proteins from a virus – binding to cell-surface receptors. A peptide will only bind if it fits like a key into a lock. |
|||
{| align="center" |
{| align="center" |
||
|| |
|| |
||
[[Image:Peptide1.PNG|center|thumb| |
[[Image:Peptide1.PNG|center|thumb|500px|A peptide (in blue) bound to a MHC Class I protein (in yellow).]] |
||
|| |
|| |
||
|
|||
|| |
|| |
||
[[Image:Peptide2.PNG|center|thumb|300px|The ends of the peptide bind inside binding pockets, while the connecting backbone extrudes above the binding groove on the surface of the MHC I molecule.]] |
|||
|} |
|} |
||
The binding is a critical step in a critical component of the immune system: it allows circulating '''T-cells to kill off infected cells.''' If this mechanism succeeds, an infection is stopped in its tracks. If it fails, then infected cells become factories for reproducing copies of the virus; full-blown disease results. Given a novel pathogen, such as SARS-Cov-2, predicting whether the immune system of an individual will do its job at fighting off the disease comes down to predicting how well the viral peptides bind to the cell-surface receptors of that person. We are tackling the problem with cloud computing resources, donated by [https://blogs.oracle.com/research/post/announcing-inaugural-cohort-oracle-research-fellows Oracle]. |
|||
'''Machine Learning Predictions''' |
|||
Predicting peptide-MHC binding is of significance in determining whether a given person's immune system can detect and effectively respond to cellular infections. [https://services.healthtech.dtu.dk/service.php?NetMHC-4.0 NetMHC] and [https://services.healthtech.dtu.dk/service.php?NetMHCpan-4.1 NetMHCpan] are state-of-the-art machine learning based tools used for this purpose. While investigating binding peptides predicted from the SARS-Cov-2 spike protein, we observed certain false positives being predicted as binders. We identified hydrophobicity as a key biochemical factor that was useful for testing the accuracy of these machine learning predictions. |
|||
{| |
{| |
||
| |
| |
||
| Line 186: | Line 178: | ||
|- valign="top" |
|- valign="top" |
||
| width="100" | '''title''': |
| width="100" | '''title''': |
||
| width="500" | [https://mriedel.ece.umn.edu/wiki/images/2/2d/Atkins_Solanki_Vasmatzis_Cornette_Riedel_Geographically_Biased_Composition_of_NetMHCpan_Training_Datasets_and_Evaluation_of_MHC-Peptide_Binding_Prediction_Accuracy_on_Novel_Alleles.pdf Geographically Biased Composition of NetMHCpan Training Datasets and Evaluation of MHC-Peptide] |
|||
| width="500" | [//http://mriedel.ece.umn.edu/wiki/images/e/e6/ISMCO_netMHC.pdf The Role of Hydrophobicity in Peptide-MHC Binding] |
|||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [[Arnav Solanki]], [ |
| Thomas Atkins, [[Arnav Solanki]], [https://www.mayo.edu/research/faculty/vasmatzis-george-ph-d/bio-00027811 George Vasmatzis], [https://orion.math.iastate.edu/cornette/ James Cornette], and [[Marc Riedel]] |
||
|- valign="top" |
|- valign="top" |
||
| ''' |
| '''appeared in''': |
||
| [https://www.frontiersin.org/journals/immunology Frontiers in Immunology], Vol. 14, 2023 |
|||
| [http://ismco.net/ 3rd International Symposium on Mathematical and Computational Oncology], 2021 |
|||
|} |
|} |
||
| align="center" width="70" | |
| align="center" width="70" | |
||
<span class="plainlinks"> |
<span class="plainlinks"> |
||
[ |
[https://www.frontiersin.org/articles/10.3389/fimmu.2023.1288105/full http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
<br> |
<br> |
||
[https://www.frontiersin.org/articles/10.3389/fimmu.2023.1288105/full Paper] |
|||
[//http://mriedel.ece.umn.edu/wiki/images/e/e6/ISMCO_netMHC.pdf Paper] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://mriedel.ece.umn.edu/wiki/images/8/8b/ISMCO2021.pdf http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> |
|||
[http://mriedel.ece.umn.edu/wiki/images/8/8b/ISMCO2021.pdf Slides] |
|||
|} |
|} |
||
'''Mechanistic Simulations''' |
|||
We are currently building a FORTRAN based simulation tool that will model peptide and MHC Class I binding mechanistically, i.e. we will incorporate several biochemical factors pertinent to binding, such as hydrophobicity, van der Waals forces, electrostatic forces, pi-interactions, etc. This work is in collaboration with the Mayo Clinic. |
|||
==Computing with Nanoscale Lattices== |
|||
"''Listen to the technology; find out what it’s telling you.''” –– '''Carver Mead''' (1934– ) |
|||
In his seminal Master's Thesis, [http://en.wikipedia.org/wiki/Claude_Shannon Claude Shannon] made the connection between Boolean algebra and switching circuits. He considered '''two-terminal''' switches corresponding to electromagnetic relays. A Boolean function can be implemented in terms of connectivity across a network of switches, often arranged in a series/parallel configuration. We have developed a method for synthesizing Boolean functions with networks of '''four-terminal switches'''. Our model is applicable for variety of nanoscale technologies, such as [http://www.sciencemag.org/content/302/5649/1377.short nanowire crossbar arrays], as [http://en.wikipedia.org/wiki/Molecular_switch molecular switch-based structures]. |
|||
{| align="center" |
|||
| |
|||
[[Image:two-terminal.gif|center|thumb|none|315px|Shannon's model: '''two-terminal switches'''. Each switch is either ON (closed) or OFF (open). A Boolean function is implemented in terms of connectivity across a network of switches, between the source S and the drain D.]] |
|||
|| |
|||
|| [[Image:four-terminal.gif|center|thumb|none|315px|Our model: '''four-terminal switches'''. Each switch is either mutually connected to its neighbors (ON) or disconnected (OFF). A Boolean function is implemented in terms of connectivity between the top and bottom plates. This network implements the same function as the two-terminal network on the left.]] |
|||
|} |
|||
{| |
{| |
||
| |
| |
||
{| style="background:#F0E68C" |
{| style="background:#F0E68C" |
||
|- valign="top" |
|- valign="top" |
||
| width="100" |'''title''': |
| width="100" | '''title''': |
||
| width="500" | [[Media:Evasive_Spike_Variants_Elucidate_the_Preservation_of_T_Cell_Immune_Response_to_the_SARS-CoV-2_Omicron_Variant.pdf|Evasive Spike Variants Elucidate the Preservation of T cell Immune Response to the SARS-CoV-2 Omicron Variant]] |
|||
| width="500" |[[Media:Altun_Riedel_Logic_Synthesis_for_Switching_Lattices.pdf | Logic Synthesis for Switching Lattices]] |
|||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [[Arnav Solanki]], [[Marc Riedel]], [https://orion.math.iastate.edu/cornette/ James Cornette], [[Julia Udell]], and [https://www.mayo.edu/research/faculty/vasmatzis-george-ph-d/bio-00027811 George Vasmatzis] |
|||
| [[Mustafa Altun]] and [[Marc Riedel]] |
|||
|- valign="top" |
|- valign="top" |
||
| ''' |
| '''appeared in''': |
||
| [https://ieeexplore.ieee.org/document/10457533 IEEE/ACM Trans. on Computational Biology and Bioinformatics], pp. 1–11, March, 2024 |
|||
| [http://www.computer.org/portal/web/tc IEEE Transactions on Computers], 2011. |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.dac.com/47th/index.aspx Design Automation Conference], Anaheim, CA, 2010. |
|||
|} |
|} |
||
| |
| width="70" align="center" | |
||
<span class="plainlinks"> |
<span class="plainlinks"> |
||
[ |
[https://mriedel.ece.umn.edu/wiki/images/1/12/Evasive_Spike_Variants_Elucidate_the_Preservation_of_T_Cell_Immune_Response_to_the_SARS-CoV-2_Omicron_Variant.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
<br> |
<br> |
||
[[Media:Evasive_Spike_Variants_Elucidate_the_Preservation_of_T_Cell_Immune_Response_to_the_SARS-CoV-2_Omicron_Variant.pdf | Paper]] |
|||
[[Media:Altun_Riedel_Logic_Synthesis_for_Switching_Lattices.pdf | Paper]] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://mriedel.ece.umn.edu/wiki/images/2/28/Altun_Riedel_Lattice-Based_Computation_of_Boolean_Functions.ppt http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/2/28/Altun_Riedel_Lattice-Based_Computation_of_Boolean_Functions.ppt Slides] |
|||
|} |
|} |
||
The impetus for nanowire-based technology is the potential density, scalability and manufacturability. Many other novel and emerging technologies fit the general model of four-terminal switches. For instance, researchers are investigating [http://en.wikipedia.org/wiki/Spin_wave spin waves]. A common feature of many emerging technologies for switching networks is that they exhibit high defect rates. |
|||
{| align="center" |
|||
| |
|||
[[Image:nano-crossbar.gif|center|thumb|315px|A nanowire crossbar switch. |
|||
The connections between horizontal and vertical |
|||
wires are FET-like junctions. When high or low voltages are applied to input nanowires, the |
|||
FET-like junctions that cross these develop a high or low impedance, respectively. |
|||
]] |
|||
|| |
|||
|| [[Image:percolation.gif|center|thumb|315px| In a switching network with defects, percolation can be exploited to produce robust Boolean functionality. Unless the defect rate exceeds an error margin, with high probability no connection forms between the top and bottom plates for logical zero ("OFF"); with high probability, a connection forms for logical one ("ON").]] |
|||
|} |
|||
We have devised a novel framework for digital computation with lattices of nanoscale switches with high defect rates, based on the mathematical phenomenon of [http://en.wikipedia.org/wiki/Percolation_theory percolation]. With random connectivity, percolation gives rise to a sharp non-linearity in the probability of global connectivity as a function of the probability of local connectivity. We exploit this phenomenon to compute Boolean functions robustly in the presence of defects. |
|||
{| |
|||
| |
|||
{| style="background:#F0E68C" |
|||
|- valign="top" |
|||
| width="100" |'''title''': |
|||
| width="500" |[[Media:Altun_Riedel_Robust_Computation_through_Percolation_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf | Synthesizing Logic with Percolation in Nanoscale Lattices]] |
|||
|- valign="top" |
|||
| '''authors''': |
|||
| [[Mustafa Altun]] and [[Marc Riedel]] |
|||
|- valign="top" |
|||
| '''appeared in''': |
|||
| [http://www.igi-global.com/Bookstore/TitleDetails.aspx?TitleId=1117&DetailsType=Description/ International Journal of Nanotechnology and Molecular Computation], <br>Vol. 3, No. 2, pp. 12–30, 2011. |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.dac.com/46th/index.aspx Design Automation Conference], San Francisco, CA, 2009. |
|||
|} |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://cadbio.com/wiki/images/3/3b/Altun_Riedel_Robust_Computation_through_Percolation_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> |
|||
[[Media:Altun_Riedel_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf | Paper]] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks">[http://mriedel.ece.umn.edu/wiki/images/f/fe/Altun_Riedel_Neuhauser_Nanoscale_Digital_Computation_Through_Percolation.ppt http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/f/fe/Altun_Riedel_Neuhauser_Nanoscale_Digital_Computation_Through_Percolation.ppt Slides] |
|||
|} |
|||
<!-- [[Image:Lattice-defects-percolation.gif|center|thumb|none|700px|In a switching network with defects, percolation can be exploited to produce robust Boolean functionality. Unless the defect rate exceeds an error margin, with high probability, no connection forms between the top and bottom plates for logical zero ("OFF"); with high probability, a connection forms for logical one ("ON").]] --> |
|||
Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
|||
==Computing with Random Bit Streams== |
==Computing with Random Bit Streams== |
||
| Line 294: | Line 217: | ||
"''To invent, all you need is a pile of junk and a good imagination.''" –– '''Thomas A. Edison''' (1847–1931) |
"''To invent, all you need is a pile of junk and a good imagination.''" –– '''Thomas A. Edison''' (1847–1931) |
||
Humans are accustomed to counting in a positional number system – [http://en.wikipedia.org/wiki/Decimal decimal] radix. Nearly all computer systems operate on another |
Humans are accustomed to counting in a positional number system – [http://en.wikipedia.org/wiki/Decimal decimal] radix. Nearly all computer systems operate on another – [http://en.wikipedia.org/wiki/Binary_numeral_system binary] radix. We are so accustomed to these systems that it counterintuitive to ask: ''can we compute using a different representatio''n? and ''why would we want to''? |
||
==== Logic |
==== Stochastic Logic ==== |
||
We advocate an alternative representation: random bit streams where the signal value is encoded by the probability of obtaining a one versus a zero. |
We advocate an alternative representation: computing on random bit streams, where the signal value is encoded by the probability of obtaining a one versus a zero. Why compute this way? Using '''stochastic logic''', we can compute complex functions with very, very simple circuits. For instance, we can perform multiplication with a single AND gate and addition with a single MUX: |
||
{| align="center" |
{| align="center" |
||
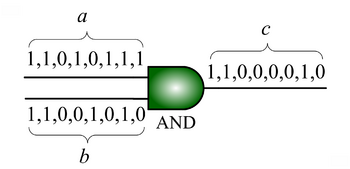
| [[Image:stochastic-multiplier.png|center|thumb|350px|'''Multiplication''' with an AND gate. Here the variables represents the ''probabilities'' of obtaining a 1 versus a 0 in stochastic bit streams. The AND gate produces an output probability |
| [[Image:stochastic-multiplier.png|center|thumb|350px|'''Multiplication''' with an AND gate. Here the variables represents the ''probabilities'' of obtaining a 1 versus a 0 in stochastic bit streams. The AND gate produces an output probability c that is the product of the of the input probabilities a and b.]] |
||
|| |
|| |
||
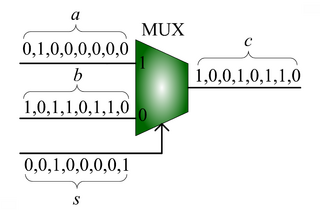
|| [[Image:stochastic-adder.png|thumb|320px|'''Scaled addition''' with a multiplexer (MUX). |
|| [[Image:stochastic-adder.png|thumb|320px|'''Scaled addition''' with a multiplexer (MUX). |
||
Given input probabilities <math>a</math>, |
Given input probabilities <math>a</math>, b and s, the MUX produces an output probability c = a s + (1 - s) b.]] |
||
|} |
|} |
||
Using conventional binary, building a circuit that computes, say a polynomial approximation to a trigonometric function such as ''tanh(x)'' or ''cos(x),'' requires ''thousands'' of logic gates. With stochastic logic, we have shown that we can compute such functions with about a ''dozen'' logic gates, so a '''100X reduction in gate count'''. Our most important contribution is a '''general methodology for synthesizing polynomial functions''' with stochastic logic, one of the seminal contributions to the field: |
|||
We have developed a general method for synthesizing digital circuitry that computes on such stochastic bit streams. Our method can be used to synthesize arbitrary polynomial functions. Through polynomial approximations, it can also be used to synthesize non-polynomial functions. Because the representation is uniform, with all bits weighted equally, the resulting circuits are highly tolerant of [http://en.wikipedia.org/wiki/Soft_error soft errors] (i.e., bit flips). |
|||
{| |
|||
| |
|||
{| style="background:#F0E68C" |
|||
|- valign="top" |
|||
| width="100" | '''title''': |
|||
| width="500" | [[Media:Qian_Riedel_Synthesizing_Logical_Computation_on_Stochastic_Bit_Streams.pdf | Synthesizing Logical Computation on Stochastic Bit Streams]] |
|||
|- valign="top" |
|||
| '''authors''': |
|||
| [[Weikang Qian]] and [[Marc Riedel]] |
|||
|- valign="top" |
|||
| '''appeared as''': |
|||
| Techincal Report, UMN |
|||
|- valign="top" |
|||
|} |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://cadbio.com/wiki/images/6/64/Qian_Riedel_Synthesizing_Logical_Computation_on_Stochastic_Bit_Streams.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> |
|||
[[Media:Qian_Riedel_Synthesizing_Logical_Computation_on_Stochastic_Bit_Streams.pdf | Paper]] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks">[http://mriedel.ece.umn.edu/files/Riedel_Synthesizing_Logical_Computation_on_Stochastic_Bit_Streams.pptx http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br>[http://mriedel.ece.umn.edu/files/Riedel_Synthesizing_Logical_Computation_on_Stochastic_Bit_Streams.pptx Slides] |
|||
|} |
|||
{| |
{| |
||
| |
| |
||
| Line 336: | Line 237: | ||
|- valign="top" |
|- valign="top" |
||
| width="100" | '''title''': |
| width="100" | '''title''': |
||
| width="500" | [//mriedel.ece.umn.edu/wiki/images/2/21/Qian_Li_Riedel_Bazargan_Lilja_An_Architecture_for_Fault-Tolerant_Computation_with_Stochastic_Logic.pdf An Architecture for Fault-Tolerant Computation with Stochastic Logic] |
|||
| width="500" | [[Media:Li_Lilja_Qian_Riedel_Bazargan_Logical_Computation_on_Stochastic_Bit_Streams_with_Linear_Finite_State_Machines.pdf | Logical Computation on Stochastic Bit Streams with Linear Finite State Machines]] |
|||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [ |
| [[Weikang Qian]], [http://www.arctic.umn.edu/people.shtml Xin Li], [[Marc Riedel]], [http://www.ece.umn.edu/users/kia/ Kia Bazargan], and [http://www.arctic.umn.edu/lilja.shtml David Lilja] |
||
|- valign="top" |
|- valign="top" |
||
| '''appeared in''': |
| '''appeared in''': |
||
| [http://ieeexplore.ieee.org/ |
| [http://ieeexplore.ieee.org/xpl/freeabs_all.jsp?arnumber=5601694 IEEE Transactions on Computers], Vol. 60, No. 1, pp. 93–105, 2011 |
||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://ieeexplore.ieee.org/xpls/abs_all.jsp?arnumber=6165056 IEEE/ACM Asia and South Pacific Design Automation Conference],<br>Sydney, Australia, 2012 |
|||
|} |
|} |
||
| |
| width="70" align="center" | |
||
<span class="plainlinks"> |
<span class="plainlinks"> |
||
[http:// |
[http://cadbio.com/wiki/images/2/21/Qian_Li_Riedel_Bazargan_Lilja_An_Architecture_for_Fault-Tolerant_Computation_with_Stochastic_Logic.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
<br> |
<br> |
||
[//mriedel.ece.umn.edu/wiki/images/2/21/Qian_Li_Riedel_Bazargan_Lilja_An_Architecture_for_Fault-Tolerant_Computation_with_Stochastic_Logic.pdf Paper] |
|||
[[Media:Li_Lilja_Qian_Riedel_Bazargan_Logical_Computation_on_Stochastic_Bit_Streams_with_Linear_Finite_State_Machines.pdf | Paper]] |
|||
|} |
|} |
||
====Logic that Generates Probabilities==== |
====Logic that Generates Probabilities==== |
||
We have also shown how to synthesize logic that transforms a set of ''source'' probabilities into different ''target'' probabilities. |
|||
Schemes for probabilistic computation can exploit physical sources to generate random values in the form of bit streams. Generally, each source has a fixed bias and so provides bits that have a specific probability of being one versus zero. If many different probability values are required, it can be difficult or expensive to generate all of these directly from physical sources. In this work, we demonstrate novel techniques for synthesizing combinational logic that transforms a set of ''source'' probabilities into different ''target'' probabilities. |
|||
<!--The problem I consider is: given a set ''S'' of ''n'' probabilistic inputs with probabilities ''p''<sub>1</sub>, ..., ''p''<sub><I>n</I></sub> of being one and a target probability ''q'', how can we synthesize a combinational circuit that takes inputs from the set ''S'' and produces an output with probability ''q'' of being one?--> |
|||
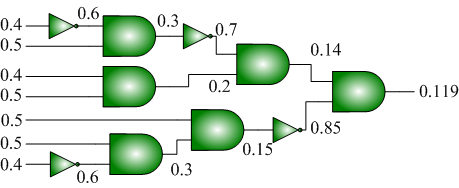
[[File:Generate_Probabilities_Example.png|center|frame|Given a set ''S'' of source probabilities {0.4, 0.5}, we can synthesize a combinational circuit to generate an arbitrary ''decimal'' output probability. The example shows how to generate 0.119. Each AND gate performs a multiplication and each inverter performs a "one-minus" operation.]] |
[[File:Generate_Probabilities_Example.png|center|frame|Given a set ''S'' of source probabilities {0.4, 0.5}, we can synthesize a combinational circuit to generate an arbitrary ''decimal'' output probability. The example shows how to generate 0.119. Each AND gate performs a multiplication and each inverter performs a "one-minus" operation.]] |
||
| Line 387: | Line 283: | ||
|} |
|} |
||
'''<big>A Deterministic Approach</big>''' |
|||
====Computing with Crappy Clocks==== |
|||
Having championed stochastic logic for many years, we decided to reexamine its foundations. Why can complex functions be computed with such simple circuits when we compute on probabilities? Intuition might suggest that somehow we are harnessing deep aspects of probability theory. ''This intuition is <u>wrong</u>''. |
|||
The keys is that we operate on uniform representation rather than a positional one. We showed that '''we can compute deterministically using the same structures that we use when computing stochastically.''' ''There is no need to do anything randomly!'' This upended the field that we had pioneered. |
|||
Clock distribution networks are a significant source of power consumption and a major design bottleneck for high-performance circuits. We have proposed a radically new approach: splitting clock domains at a very fine level, with domains consisting of only a handful of gates each. These domains are synchrnonized by "crappy clocks", generated locally with inverter rings. This is feasible if one adopts the paradigm of computing on randomized bit streams. |
|||
[[File:polysynchronous-and.png|center|frame|thumb|Stochastic multiplication using an AND gate with unsynchronized random bit streams. The stochastic paradigm can tolerate arbitrarly high clock skew. Accordingly, one can replace an expensive global clock distribution network with cheap local clocks, generated by inverter rings – "crappy clocks".]] |
|||
{| |
{| |
||
| |
| |
||
| Line 396: | Line 294: | ||
|- valign="top" |
|- valign="top" |
||
| width="100" | '''title''': |
| width="100" | '''title''': |
||
| width="500" | [//mriedel.ece.umn.edu/wiki/images/c/c9/Najafi_Jenson_Lilja_Riedel_Performing_Stochastic_Computation_Deterministically.pdf Performing Stochastic Computation Deterministically] |
|||
| width="500" | [[Media: Najafi_Lilja_Riedel_Bazargan_Polysynchronous_Stochastic_Circuits.pdf | Polysynchronous Stochastic Circuits]] |
|||
|- valign="top" |
|- valign="top" |
||
| '''authors''': |
| '''authors''': |
||
| [[M. |
| [[Devon Jenson]], [[M. Hassan Najafi]], [https://ece.umn.edu/directory/lilja-david/ David Lilja], and [[Marc Riedel]] |
||
|- valign="top" |
|- valign="top" |
||
| ''' |
| '''appeared in''': |
||
| [https://ieeexplore.ieee.org/document/8793244 IEEE Trans. on Very Large Scale Integration Systems],<br>Vol. 27, No. 29, pp. 2925–2938, 2019 |
|||
| [http://www.amsv.umac.mo/aspdac2016/ IEEE/ACM Asia and South Pacific Design Automation Conference], 2016 |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [https://iscas2020.org IEEE International Symposium of Circuits and Systems], 2020 |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.ieee.org/conferences_events/conferences/conferencedetails/index.html?Conf_ID=38629 IEEE/ACM International Conference on Computer-Aided Design], 2016 |
|||
|} |
|} |
||
| |
| width="70" align="center" | |
||
<span class="plainlinks"> |
<span class="plainlinks"> |
||
[http:// |
[http://mriedel.ece.umn.edu/wiki/images/c/c9/Najafi_Jenson_Lilja_Riedel_Performing_Stochastic_Computation_Deterministically.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
||
<br> |
<br> |
||
[//mriedel.ece.umn.edu/wiki/images/c/c9/Najafi_Jenson_Lilja_Riedel_Performing_Stochastic_Computation_Deterministically.pdf Paper] |
|||
[[Media:Najafi_Lilja_Riedel_Bazargan_Polysynchronous_Stochastic_Circuits.pdf | Paper]] |
|||
| width="70" align="center" | |
|||
<span class="plainlinks">[http://cadbio.com/wiki/images/4/4f/Jenson_Riedel_A_Deterministic_Approach_to_Stochastic_Computing.pptx http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br>[http://cadbio.com/wiki/images/4/4f/Jenson_Riedel_A_Deterministic_Approach_to_Stochastic_Computing.pptx Slides] |
|||
|} |
|} |
||
====Time-Encoded Computing==== |
|||
Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
|||
Computing deterministically on bit streams really means that, instead of encoding data in '''''space''''', we encode them '''''time'''''. The time-encoding consists of periodic signals, with the value encoded as the fraction of the time that the signal is in the high (on) state compared to the low (off) state in each cycle. |
|||
{| align="center" |
|||
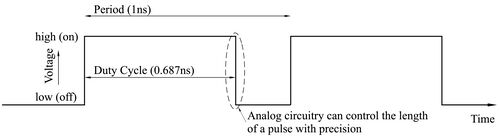
|[[File:Analog-in-time.jpg|center|thumb|500px|Encoding a value in time. The value represented is the fraction of the time that the signal is high in each cycle, in this case 0.687.]] |
|||
| |
|||
||[[File:Multiplicaiton-on-time-encoding-signals.jpg|thumb|500px|Multiplication with a single AND gate, operating on deterministic periodic signals. ]] |
|||
|} |
|||
As technology has scaled and device sizes have gotten smaller, the supply voltages have dropped while the device speeds have improved. Control of the dynamic range in the voltage domain is limited; however, control of the length of pulses in the time domain can be precise. Encoding data in the time domain can be done more accurately and more efficiently than converting signals into binary radix. So we can compute ''more precisely,'' ''faster'', and with ''fewer logic gates:'' |
|||
{| |
|||
|- |
|||
| |
|||
{| style="background:#F0E68C" |
|||
|- valign="top" |
|||
| width="100" | '''title''': |
|||
| width="500" | [//mriedel.ece.umn.edu/wiki/images/b/b0/Najafi_Jamali_Zavareh_Lilja_Riedel_Bazargan_Harjani_Time_Encoded_Values_for_Highly_Efficient_Stochastic_Circuits.pdf Time-Encoded Values for Highly Efficient Stochastic Circuits] |
|||
|- valign="top" |
|||
| '''authors''': |
|||
| [[M. Hassan Najafi]], S. Jamali-Zavareh, [http://www.arctic.umn.edu/lilja.shtml David Lilja], [[Marc Riedel]], [http://people.ece.umn.edu/~kia Kia Bazargan] and<br>[http://people.ece.umn.edu/~harjani/ Ramesh Harjani] |
|||
|- valign="top" |
|||
| '''appeared in''': |
|||
| [http://ieeexplore.ieee.org/xpl/aboutJournal.jsp?punumber=92 IEEE Trans. on Very Large Scale Integration Systems],<br>Vol. 25, No. 5, pp. 1644–1657, 2017 |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.ieee.org/conferences_events/conferences/conferencedetails/index.html?Conf_ID=33621 IEEE International Symposium on Circuits and Systems], 2017 |
|||
|} |
|||
| width="70" align="center" | |
|||
<span class="plainlinks"> |
|||
[http://www.mriedel.ece.umn.edu/wiki/images/b/b0/Najafi_Jamali_Zavareh_Lilja_Riedel_Bazargan_Harjani_Time_Encoded_Values_for_Highly_Efficient_Stochastic_Circuits.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> |
|||
[//mriedel.ece.umn.edu/wiki/images/b/b0/Najafi_Jamali_Zavareh_Lilja_Riedel_Bazargan_Harjani_Time_Encoded_Values_for_Highly_Efficient_Stochastic_Circuits.pdf Paper] |
|||
|}Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
|||
==Computing with Feedback== |
==Computing with Feedback== |
||
| Line 456: | Line 395: | ||
|} |
|} |
||
Please see our [[Papers,_Theses,_and_Presentations | Publications]] page for more of our papers on this topic. |
Please see our [[Papers,_Theses,_and_Presentations | Publications]] page for more of our papers on this topic. |
||
==Computing with Nanoscale Lattices== |
|||
"''Listen to the technology; find out what it’s telling you.''” –– '''Carver Mead''' (1934– ) |
|||
In his seminal Master's Thesis, [http://en.wikipedia.org/wiki/Claude_Shannon Claude Shannon] made the connection between Boolean algebra and switching circuits. He considered '''two-terminal''' switches corresponding to electromagnetic relays. A Boolean function can be implemented in terms of connectivity across a network of switches, often arranged in a series/parallel configuration. We have developed a method for synthesizing Boolean functions with networks of '''four-terminal switches'''. Our model is applicable for variety of nanoscale technologies, such as [http://www.sciencemag.org/content/302/5649/1377.short nanowire crossbar arrays], as [http://en.wikipedia.org/wiki/Molecular_switch molecular switch-based structures]. |
|||
{| align="center" |
|||
| |
|||
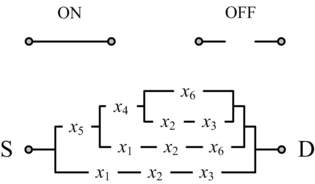
[[Image:two-terminal.gif|center|thumb|none|315px|Shannon's model: '''two-terminal switches'''. Each switch is either ON (closed) or OFF (open). A Boolean function is implemented in terms of connectivity across a network of switches, between the source S and the drain D.]] |
|||
|| |
|||
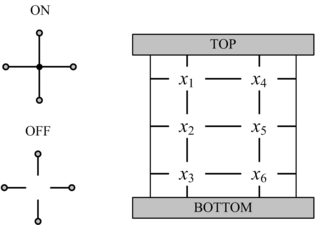
|| [[Image:four-terminal.gif|center|thumb|none|315px|Our model: '''four-terminal switches'''. Each switch is either mutually connected to its neighbors (ON) or disconnected (OFF). A Boolean function is implemented in terms of connectivity between the top and bottom plates. This network implements the same function as the two-terminal network on the left.]] |
|||
|} |
|||
{| |
|||
| |
|||
{| style="background:#F0E68C" |
|||
|- valign="top" |
|||
| width="100" |'''title''': |
|||
| width="500" |[[Media:Altun_Riedel_Logic_Synthesis_for_Switching_Lattices.pdf | Logic Synthesis for Switching Lattices]] |
|||
|- valign="top" |
|||
| '''authors''': |
|||
| [[Mustafa Altun]] and [[Marc Riedel]] |
|||
|- valign="top" |
|||
| '''will appear in''': |
|||
| [http://www.computer.org/portal/web/tc IEEE Transactions on Computers], 2011. |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.dac.com/47th/index.aspx Design Automation Conference], Anaheim, CA, 2010. |
|||
|} |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://cadbio.com/wiki/images/c/ca/Altun_Riedel_Logic_Synthesis_for_Switching_Lattices.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> |
|||
[[Media:Altun_Riedel_Logic_Synthesis_for_Switching_Lattices.pdf | Paper]] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://mriedel.ece.umn.edu/wiki/images/2/28/Altun_Riedel_Lattice-Based_Computation_of_Boolean_Functions.ppt http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/2/28/Altun_Riedel_Lattice-Based_Computation_of_Boolean_Functions.ppt Slides] |
|||
|} |
|||
The impetus for nanowire-based technology is the potential density, scalability and manufacturability. Many other novel and emerging technologies fit the general model of four-terminal switches. For instance, researchers are investigating [http://en.wikipedia.org/wiki/Spin_wave spin waves]. A common feature of many emerging technologies for switching networks is that they exhibit high defect rates. |
|||
{| align="center" |
|||
| |
|||
[[Image:nano-crossbar.gif|center|thumb|315px|A nanowire crossbar switch. |
|||
The connections between horizontal and vertical |
|||
wires are FET-like junctions. When high or low voltages are applied to input nanowires, the |
|||
FET-like junctions that cross these develop a high or low impedance, respectively. |
|||
]] |
|||
|| |
|||
|| [[Image:percolation.gif|center|thumb|260.994x260.994px| In a switching network with defects, percolation can be exploited to produce robust Boolean functionality. ]] |
|||
|} |
|||
We have devised a novel framework for digital computation with lattices of nanoscale switches with high defect rates, based on the mathematical phenomenon of [http://en.wikipedia.org/wiki/Percolation_theory percolation]. With random connectivity, percolation gives rise to a sharp non-linearity in the probability of global connectivity as a function of the probability of local connectivity. We exploit this phenomenon to compute Boolean functions robustly in the presence of defects. |
|||
{| |
|||
| |
|||
{| style="background:#F0E68C" |
|||
|- valign="top" |
|||
| width="100" |'''title''': |
|||
| width="500" |[[Media:Altun_Riedel_Robust_Computation_through_Percolation_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf | Synthesizing Logic with Percolation in Nanoscale Lattices]] |
|||
|- valign="top" |
|||
| '''authors''': |
|||
| [[Mustafa Altun]] and [[Marc Riedel]] |
|||
|- valign="top" |
|||
| '''appeared in''': |
|||
| [http://www.igi-global.com/Bookstore/TitleDetails.aspx?TitleId=1117&DetailsType=Description/ International Journal of Nanotechnology and Molecular Computation], <br>Vol. 3, No. 2, pp. 12–30, 2011. |
|||
|- valign="top" |
|||
| '''presented at''': |
|||
| [http://www.dac.com/46th/index.aspx Design Automation Conference], San Francisco, CA, 2009. |
|||
|} |
|||
| align="center" width="70" | |
|||
<span class="plainlinks"> |
|||
[http://cadbio.com/wiki/images/3/3b/Altun_Riedel_Robust_Computation_through_Percolation_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf http://mriedel.ece.umn.edu/wiki/images/0/04/Pdf.jpg]</span> |
|||
<br> |
|||
[[Media:Altun_Riedel_Synthesizing_Logic_with_Percolation_in_Nanoscale_Lattices.pdf | Paper]] |
|||
| align="center" width="70" | |
|||
<span class="plainlinks">[http://mriedel.ece.umn.edu/wiki/images/f/fe/Altun_Riedel_Neuhauser_Nanoscale_Digital_Computation_Through_Percolation.ppt http://mriedel.ece.umn.edu/wiki/images/3/36/Ppt.jpg]</span> |
|||
<br> [http://mriedel.ece.umn.edu/wiki/images/f/fe/Altun_Riedel_Neuhauser_Nanoscale_Digital_Computation_Through_Percolation.ppt Slides] |
|||
|} |
|||
Please see our "[[Papers,_Theses,_and_Presentations |Publications]]" page for more of our papers on these topics. |
|||
==Algorithms and Data Structures== |
==Algorithms and Data Structures== |
||
| Line 461: | Line 478: | ||
"''There are two kinds of people in the world: those who divide the world into two kinds of people, and those who don't.''" –– '''Robert Charles Benchley''' (1889–1945) |
"''There are two kinds of people in the world: those who divide the world into two kinds of people, and those who don't.''" –– '''Robert Charles Benchley''' (1889–1945) |
||
Consider the task of designing a digital circuit with 256 inputs. From a mathematical standpoint, such a circuit performs mappings from a space of <math>2^{256}</math> Boolean input values to Boolean output values. (The number of rows in a [http://en.wikipedia.org/wiki/Truth_table truth table] for such a function is approximately equal to [http://en.wikipedia.org/wiki/Observable_universe#Matter_content the number of atoms in the universe] – <math>10^{77}</math> rows versus <math>10^{79}</math> atoms!) Verifying such a function, let alone designing the corresponding circuit, would seem to be an intractable problem. |
Consider the task of designing a digital circuit with 256 inputs. From a mathematical standpoint, such a circuit performs mappings from a space of <math>2^{256}</math> Boolean input values to Boolean output values. (The number of rows in a [http://en.wikipedia.org/wiki/Truth_table truth table] for such a function is approximately equal to [http://en.wikipedia.org/wiki/Observable_universe#Matter_content the number of atoms in the universe] – <math>10^{77}</math> rows versus <math>10^{79}</math> atoms!) Verifying such a function, let alone designing the corresponding circuit, would seem to be an intractable problem. |
||
Circuit designers have succeeded in their endeavor largely as a result of innovations in the data structures and algorithms used to represent and manipulate [http://en.wikipedia.org/wiki/Boolean_function Boolean functions]. We have developed novel, efficient techniques for synthesizing functional dependencies based on so-called [http://en.wikipedia.org/wiki/Boolean_satisfiability_problem SAT-solving algorithms]. We use [http://en.wikipedia.org/wiki/Craig_interpolation Craig Interpolation] to generate circuits from the corresponding Boolean functions. |
|||
{| align="center" |
{| align="center" |
||
| [[Image:sat-squid.jpg|center|thumb|350px|A circuit construct for SAT-based verification.]] |
| [[Image:sat-squid.jpg|center|thumb|350px|A circuit construct for SAT-based verification.]] |
||
| Line 492: | Line 511: | ||
"''Mathematics may be defined as the subject in which we never know what we are talking about, nor whether what we are saying is true.''" –– '''Bertrand Russell''' (1872–1970) |
"''Mathematics may be defined as the subject in which we never know what we are talking about, nor whether what we are saying is true.''" –– '''Bertrand Russell''' (1872–1970) |
||
[[Image:xkcd_disciplines_by_purity.png|center|thumb|740x740px]] |
|||
The great mathematician [http://en.wikipedia.org/wiki/John_von_Neumann John von Neumann] articulated the view that research should never meander too far down theoretical paths; it should always be guided by potential applications. This view was not based on concerns about the relevance of his profession; rather, in his judgment, real-world applications give rise to the most ''interesting'' problems for mathematicians to tackle. At their core, most of our research contributions are mathematical contributions. The tools of our trade are [http://en.wikipedia.org/wiki/Discrete_mathematics discrete math], including [http://en.wikipedia.org/wiki/Combinatorics combinatorics] and [http://en.wikipedia.org/wiki/Probability probability theory]. |
The great mathematician [http://en.wikipedia.org/wiki/John_von_Neumann John von Neumann] articulated the view that research should never meander too far down theoretical paths; it should always be guided by potential applications. This view was not based on concerns about the relevance of his profession; rather, in his judgment, real-world applications give rise to the most ''interesting'' problems for mathematicians to tackle. At their core, most of our research contributions are mathematical contributions. The tools of our trade are [http://en.wikipedia.org/wiki/Discrete_mathematics discrete math], including [http://en.wikipedia.org/wiki/Combinatorics combinatorics] and [http://en.wikipedia.org/wiki/Probability probability theory]. |
||
[[Image:papa-handwriting.gif|center|thumb|none|550px|Mathematics, before the era of [http://en.wikipedia.org/wiki/LaTeX LaTeX].]] |
|||
{| |
{| |
||
| |
| |
||
Latest revision as of 18:03, 5 May 2026
"You see things; and you say, 'Why?' But I dream things that never were; and I say, 'Why not?'"
–– George Bernard Shaw (1856 –1950)
Our research spans different disciplines ranging from digital circuit design, to algorithms, to mathematics, to synthetic biology. It tends to be inductive (as opposed to deductive) and conceptual (as opposed to applied). A recurring theme is building systems that compute in novel or unexpected ways with new and emerging technologies.
Storing Data with Molecules
All new ideas pass through three stages:
- It can't be done.
- It probably can be done, but it's not worth doing.
- I knew it was a good idea all along!
––Arthur C. Clarke (1917–2008)
Ever since Watson and Crick first described the molecular structure of DNA, its information-bearing potential has been apparent. With each nucleotide in the sequence drawn from the four-valued alphabet of {A, T , C, G}, a molecule of DNA with n nucleotides stores 2n bits of data.
- Could we store data for our computer systems in DNA? "Can't be done – too hard."
- Is it worth doing? "Definitely not. It will never work as well as our hard drives do."
- But one can store so much data so efficiently! "I knew it was a good idea all along!"
|

|

|
Computing with Molecules
"Biology is the most powerful technology ever created. DNA is software, protein are hardware, cells are factories."
––Arvind Gupta (1953– )
Computing has escaped! It has gone from desktops and data centers into the wild. Embedded microcontrollers – found in our gadgets, our buildings, and even our bodies – are transforming our lives. And yet, there are limits to where silicon can go and where it can compute effectively. It is a foreign object that requires a electrical power source.
We are studying novel types of computing systems that are not foreign, but rather an integral part of their physical and chemical environments: systems that compute directly with molecules. A simple but radical idea: compute with acids and bases. An acidic solution corresponds to a "1" and a basic solution to "0".
|
It's more complex that acids and bases, but DNA is a terrific chassis for computing. We have developed "CS 101" algorithms with DNA: Sorting, Shifting and Searching:
|
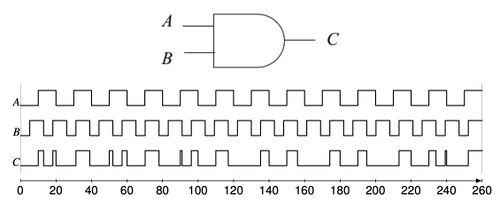
Based on a bistable mechanism for representing bits, we have implemented logic gates such AND, OR, and XOR gates, as well as sequential components such as latches and flip-flops with DNA. Using these components, we have built full-fledged digital circuits such as a binary counters and linear feedback shift registers.
|
Also, we have performed signal processing including operations such as filtering and fast-fourier transforms (FFTs) with DNA.
|
Please see our "Publications" page for more of our papers on these topics.
Computational Immunology
“Physics is the study of the simple things in the Universe. Biology is the study of the complex ones. ”
–– Richard Dawkins (1941– )
We are studying a problem that computer science currently judges to be very difficult: predicting cellular immunity. It centers on the question of how strongly molecules binds to one another. The molecules in question are peptides – fragments of proteins from a virus – binding to cell-surface receptors. A peptide will only bind if it fits like a key into a lock.
The binding is a critical step in a critical component of the immune system: it allows circulating T-cells to kill off infected cells. If this mechanism succeeds, an infection is stopped in its tracks. If it fails, then infected cells become factories for reproducing copies of the virus; full-blown disease results. Given a novel pathogen, such as SARS-Cov-2, predicting whether the immune system of an individual will do its job at fighting off the disease comes down to predicting how well the viral peptides bind to the cell-surface receptors of that person. We are tackling the problem with cloud computing resources, donated by Oracle.
|
|
Computing with Random Bit Streams
"To invent, all you need is a pile of junk and a good imagination." –– Thomas A. Edison (1847–1931)
Humans are accustomed to counting in a positional number system – decimal radix. Nearly all computer systems operate on another – binary radix. We are so accustomed to these systems that it counterintuitive to ask: can we compute using a different representation? and why would we want to?
Stochastic Logic
We advocate an alternative representation: computing on random bit streams, where the signal value is encoded by the probability of obtaining a one versus a zero. Why compute this way? Using stochastic logic, we can compute complex functions with very, very simple circuits. For instance, we can perform multiplication with a single AND gate and addition with a single MUX:
Using conventional binary, building a circuit that computes, say a polynomial approximation to a trigonometric function such as tanh(x) or cos(x), requires thousands of logic gates. With stochastic logic, we have shown that we can compute such functions with about a dozen logic gates, so a 100X reduction in gate count. Our most important contribution is a general methodology for synthesizing polynomial functions with stochastic logic, one of the seminal contributions to the field:
|
Logic that Generates Probabilities
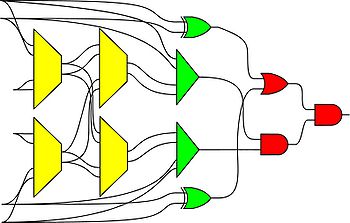
We have also shown how to synthesize logic that transforms a set of source probabilities into different target probabilities.
|
A Deterministic Approach
Having championed stochastic logic for many years, we decided to reexamine its foundations. Why can complex functions be computed with such simple circuits when we compute on probabilities? Intuition might suggest that somehow we are harnessing deep aspects of probability theory. This intuition is wrong.
The keys is that we operate on uniform representation rather than a positional one. We showed that we can compute deterministically using the same structures that we use when computing stochastically. There is no need to do anything randomly! This upended the field that we had pioneered.
|
Time-Encoded Computing
Computing deterministically on bit streams really means that, instead of encoding data in space, we encode them time. The time-encoding consists of periodic signals, with the value encoded as the fraction of the time that the signal is in the high (on) state compared to the low (off) state in each cycle.
As technology has scaled and device sizes have gotten smaller, the supply voltages have dropped while the device speeds have improved. Control of the dynamic range in the voltage domain is limited; however, control of the length of pulses in the time domain can be precise. Encoding data in the time domain can be done more accurately and more efficiently than converting signals into binary radix. So we can compute more precisely, faster, and with fewer logic gates:
|
Please see our "Publications" page for more of our papers on these topics.
Computing with Feedback
"A person with a new idea is a crank until the idea succeeds." –– Mark Twain (1835–1910)
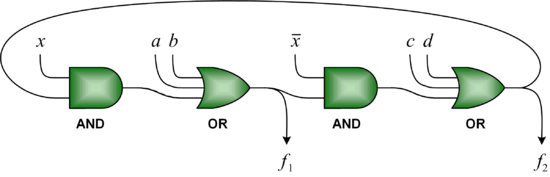
The accepted wisdom is that combinational circuits (i.e., memoryless circuits) must have acyclic (i.e., loop-free or feed-forward) topologies. And yet simple examples suggest that this need not be so. We advocate the design of cyclic combinational circuits (i.e., circuits with loops or feedback paths). We have proposed a methodology for synthesizing such circuits and demonstrated that it produces significant improvements in area and in delay.
|
Please see our Publications page for more of our papers on this topic.
Computing with Nanoscale Lattices
"Listen to the technology; find out what it’s telling you.” –– Carver Mead (1934– )
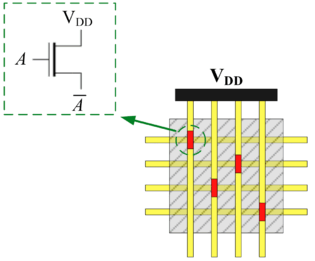
In his seminal Master's Thesis, Claude Shannon made the connection between Boolean algebra and switching circuits. He considered two-terminal switches corresponding to electromagnetic relays. A Boolean function can be implemented in terms of connectivity across a network of switches, often arranged in a series/parallel configuration. We have developed a method for synthesizing Boolean functions with networks of four-terminal switches. Our model is applicable for variety of nanoscale technologies, such as nanowire crossbar arrays, as molecular switch-based structures.
|
The impetus for nanowire-based technology is the potential density, scalability and manufacturability. Many other novel and emerging technologies fit the general model of four-terminal switches. For instance, researchers are investigating spin waves. A common feature of many emerging technologies for switching networks is that they exhibit high defect rates.
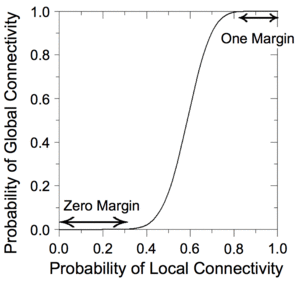
We have devised a novel framework for digital computation with lattices of nanoscale switches with high defect rates, based on the mathematical phenomenon of percolation. With random connectivity, percolation gives rise to a sharp non-linearity in the probability of global connectivity as a function of the probability of local connectivity. We exploit this phenomenon to compute Boolean functions robustly in the presence of defects.
|
Please see our "Publications" page for more of our papers on these topics.
Algorithms and Data Structures
"There are two kinds of people in the world: those who divide the world into two kinds of people, and those who don't." –– Robert Charles Benchley (1889–1945)
Consider the task of designing a digital circuit with 256 inputs. From a mathematical standpoint, such a circuit performs mappings from a space of <math>2^{256}</math> Boolean input values to Boolean output values. (The number of rows in a truth table for such a function is approximately equal to the number of atoms in the universe – <math>10^{77}</math> rows versus <math>10^{79}</math> atoms!) Verifying such a function, let alone designing the corresponding circuit, would seem to be an intractable problem.
Circuit designers have succeeded in their endeavor largely as a result of innovations in the data structures and algorithms used to represent and manipulate Boolean functions. We have developed novel, efficient techniques for synthesizing functional dependencies based on so-called SAT-solving algorithms. We use Craig Interpolation to generate circuits from the corresponding Boolean functions.
|
Please see our "Publications" page for more of our papers on this topic. (Papers on SAT-based circuit verification, that is, not on squids.)
Mathematics
"Mathematics may be defined as the subject in which we never know what we are talking about, nor whether what we are saying is true." –– Bertrand Russell (1872–1970)
The great mathematician John von Neumann articulated the view that research should never meander too far down theoretical paths; it should always be guided by potential applications. This view was not based on concerns about the relevance of his profession; rather, in his judgment, real-world applications give rise to the most interesting problems for mathematicians to tackle. At their core, most of our research contributions are mathematical contributions. The tools of our trade are discrete math, including combinatorics and probability theory.
|
Please see our "Publications" page for more of our papers on this topic.